Medicine by Alexandros G. Sfakianakis,Anapafseos 5 Agios Nikolaos 72100 Crete Greece,00302841026182,00306932607174,alsfakia@gmail.com,
Πληροφορίες
Ετικέτες
Τρίτη 7 Ιανουαρίου 2020
DNA methylation profiling in recurrent miscarriage
DNA methylation profiling in recurrent miscarriage:
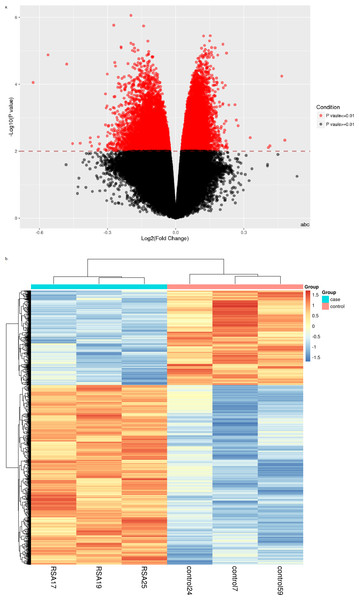
 Recurrent miscarriage (RM) is a complex clinical problem. However, specific diagnostic biomarkers and candidate regulatory targets have not yet been identified. To explore RM-related biological markers and processes, we performed a genome-wide DNA methylation analysis using the Illumina Infinium HumanMethylation450 array platform. Methylation variable positions and differentially methylated regions (DMRs) were selected using the Limma package in R language. Thereafter, gene ontology (GO) enrichment analysis and pathway enrichment analysis were performed on these DMRs. A total of 1,799 DMRs were filtered out between patients with RM and healthy pregnant women. The GO terms were mainly related to system development, plasma membrane part, and sequence-specific DNA binding, while the enriched pathways included cell adhesion molecules, type I diabetes mellitus, and ECM–receptor interactions. In addition, genes, including ABR, ALCAM, HLA-E, HLA-G, and ISG15, were obtained. These genes may be potential candidates for diagnostic biomarkers and possible regulatory targets in RM. We then detected the mRNA expression levels of the candidate genes. The mRNA expression levels of the candidate genes in the RM group were significantly higher than those in the control group. However, additional research is still required to confirm their potential roles in the occurrence of RM.
Recurrent miscarriage (RM) is a complex clinical problem. However, specific diagnostic biomarkers and candidate regulatory targets have not yet been identified. To explore RM-related biological markers and processes, we performed a genome-wide DNA methylation analysis using the Illumina Infinium HumanMethylation450 array platform. Methylation variable positions and differentially methylated regions (DMRs) were selected using the Limma package in R language. Thereafter, gene ontology (GO) enrichment analysis and pathway enrichment analysis were performed on these DMRs. A total of 1,799 DMRs were filtered out between patients with RM and healthy pregnant women. The GO terms were mainly related to system development, plasma membrane part, and sequence-specific DNA binding, while the enriched pathways included cell adhesion molecules, type I diabetes mellitus, and ECM–receptor interactions. In addition, genes, including ABR, ALCAM, HLA-E, HLA-G, and ISG15, were obtained. These genes may be potential candidates for diagnostic biomarkers and possible regulatory targets in RM. We then detected the mRNA expression levels of the candidate genes. The mRNA expression levels of the candidate genes in the RM group were significantly higher than those in the control group. However, additional research is still required to confirm their potential roles in the occurrence of RM.

Αναρτήθηκε από
Medicine by Alexandros G. Sfakianakis,Anapafseos 5 Agios Nikolaos 72100 Crete Greece,00302841026182,00306932607174,alsfakia@gmail.com,
στις
1:45 π.μ.

Ετικέτες
00302841026182,
00306932607174,
alsfakia@gmail.com,
Anapafseos 5 Agios Nikolaos 72100 Crete Greece,
Medicine by Alexandros G. Sfakianakis,
Telephone consultation 11855 int 1193
Εγγραφή σε:
Σχόλια ανάρτησης (Atom)
Αρχειοθήκη ιστολογίου
-
►
2023
(276)
- ► Φεβρουαρίου (133)
- ► Ιανουαρίου (143)
-
►
2022
(1976)
- ► Δεκεμβρίου (116)
- ► Σεπτεμβρίου (158)
- ► Φεβρουαρίου (165)
- ► Ιανουαρίου (161)
-
►
2021
(3661)
- ► Δεκεμβρίου (161)
- ► Σεπτεμβρίου (274)
- ► Φεβρουαρίου (64)
- ► Ιανουαρίου (368)
-
▼
2020
(4554)
- ► Δεκεμβρίου (400)
- ► Σεπτεμβρίου (381)
- ► Φεβρουαρίου (638)
-
▼
Ιανουαρίου
(691)
- Medicine by Alexandros G. Sfakianakis,Anapafseos 5...
- 49 journals
- MDPI
- MDPI
- Antioxidant potential of Sargassum horneri extract...
- Tuberculous Retropharyngeal Abscess
- Opioids and Cannabinoids for Osteoarthritis: Eithe...
- Risk factors for sepsis and associated mortality a...
- Microbiome Composition in Pediatric Populations fr...
- In vivo MRS measurement of 2‐hydroxyglutarate in p...
- Thirty-Day Hospital Readmissions in a Care Transit...
- Mammary Extranodal Rosai-Dorfman Disease With and ...
- Oral health-related quality of life in tumour pati...
- Regulation of immune cell metabolism by cancer cel...
- Recurrent RET Gene Fusions in Pediatric Spindle Me...
- Elevated levels of tumor apolipoprotein-D independ...
- Digital pathology for primary diagnosis of screen-...
- Aberrant von Willebrand Factor Expression of Sinus...
- Molecular basis and restoration of function defici...
- Evolutionary repression of chondrogenic genes in t...
- Protein scaffold involving MSMEG_1285 maintain cel...
- Human proteinase 3 resistance to inhibition extend...
- The role of regulatory T cells in allergic rhiniti...
- Videolaryngoscope-assisted coblation of epiglottic...
- Mid-term evaluation of perioperative i.v. corticos...
- Lymph node density as a predictive factor for wors...
- Intravenous immunoglobulin for suspected or proven...
- Intravenous immunoglobulin for preventing infectio...
- Baricitinib in Patients with Moderate-to-Severe At...
- The life cycle of cancer-associated fibroblasts wi...
- High Neutrophil Count as a Negative Prognostic Fac...
- Age-related structural alterations of skeletal mus...
- Printability of pulp derived crystal, fibril and b...
- Nasal septum deviation can be a normal variation a...
- New features in the management of labio-maxillo-pa...
- Psychological morbidity among forcibly displaced c...
- Effects of chronic low-dose aspirin treatment on t...
- Thrombosis among patients with JAK2V617F -mutated ...
- Economics of alternative dosing strategies for pem...
- Septicemia after chemotherapy for childhood acute ...
- Detection of perilymphatic fistula in labyrinthine...
- Role of the human papillomavirus in malignant tran...
- The Protective Effect of Echinochrome A on Extrace...
- Tools for Analysis of the Microbiome
- Retrograde Transvenous Obliteration (RTO): A New T...
- Diet and Gut Microbes Act Coordinately to Enhance ...
- Microbial-Based and Microbial-Targeted Therapies f...
- Theory and applications of differential scanning f...
- Design and Synthesis of Arf1-Targeting γ-Dipeptide...
- Asiatic Acid, Extracted from Centella asiatica and...
- Calcium Electroporation -A Novel Cancer Treatment ...
- Detection of early adenocarcinoma of the esophagog...
- Deaf patients with bi-allelic and mono-allelic GJB...
- Hypoglossal nerve stimulation therapy does not alt...
- White matter alteration and autonomic impairment i...
- Study Design Considerations for Sleep Disordered B...
- Utility of the modified Mallampati grade and Fried...
- The detection of Cryptococcus in skeletal infectio...
- Developing a virus-microRNA interactome using cyto...
- The Influence of ACE Inhibition on C1-Inhibitor: A...
- Cranio-Corpo-Graphy (CCG) is a medical investigati...
- Usefulness of Ultrasound-Computer-Craniocorpograph...
- Mid-term evaluation of perioperative i.v. corticos...
- Lymph node density as a predictive factor for wors...
- HPV16 is sufficient to induce squamous cell carcin...
- Stathmin guides personalized therapy in oral squam...
- Histological characteristics of early-stage oral t...
- Multi-Institutional Regional Otolaryngology Bootcamp.
- Are Children Scheduled for Ventilation Tubes Inser...
- Cerumen Management: An Updated Clinical Review and...
- Pou3f4-Expressing Otic Mesenchyme Cells Promote Sp...
- Antibiotic prescription rate for upper respiratory...
- Effect of Poguntano leaves extract (Picria fel-ter...
- Presentation of Infantile Hemangiopericytoma/Solit...
- Elucidating Molecular Interactions of Ten Natural ...
- Correlation between work impairment, scores of rhi...
- Study of the Expression Transition of Cardiac Myos...
- Medicine by Alexandros G. Sfakianakis,Anapafseos 5...
- Medicine by Alexandros G. Sfakianakis,Anapafseos 5...
- 63 JOURNALS
- MDPI
- MDPI
- MDPI
- Trends in nutritional intake and coronary risk fac...
- Efficacy and safety of interleukin‐17 inhibitors f...
- Identification of functional domains of the minor ...
- miR-331-3p Inhibits Proliferation and Promotes Apo...
- Teleassessment of Gait and Gait Aids: Validity and...
- Piezosurgery in Translabyrinthine-Approach Exposur...
- Performing MRI Scans on Cochlear Implant and Audit...
- High-Dose Furosemide Enhances the Magnetic Resonan...
- Anatomical and Functional Characterization in Chil...
- Metallothioneins conferring cardioprotection in pu...
- Reducing the progression of Alzheimer's disease in...
- Enhanced topical cutaneous delivery of indocyanine...
- Contrast-Enhanced Ultrasound-Guided Percutaneous B...
- Diagnosis of Acute Heart Failure Using Inferior Ve...
- The modified bipedicled flap for reconstruction of...
- Correlation between superficial and deep lymphatic...
- Immunological response of allogeneic bone grafting...

Δεν υπάρχουν σχόλια:
Δημοσίευση σχολίου